
Quick start
Introduction
Welcome to WIMM Centre for Computational Biology (CCB). If you find an error in any of these pages or there's information which isn't provided that you feel would be useful, please contact us via help@imm.ox.ac.uk. Our documentation is important to us and we treat errors and omissions as problems to be fixed.
Policies
In using our systems, you are agreeing to our terms & conditions and acceptable-use. If you wish to withdraw your agreement at any time, please inform us so that we can remove your access for you.
Getting help
Commonly asked questions are answered in our FAQ. Otherwise, you can email the CCB team at help@imm.ox.ac.uk. Using this address ensures your query enters our support system. This means it is assigned a tracking number, and the appropriate member(s) of our team will pick it up. Avoid directly mailing a team member, as your email is likely to be lost amidst existing traffic and take longer to answer. We also recommend that you send a single question or request per email, and that you always send new requests as new emails and not as replies to old ones. Doing this increases likelihood of a quick answer, and reduces likelihood of us thinking request already fulfilled.
Applying for access
To find out how to apply for a CCB account or to get storage space, please see accounts.
Differences between JADE and CentOS 7
If you're an existing user there are some important differences between JADE and CentOS 7. You can read more about these in differences.
Logging in
-
You must be connecting from an Oxford University network to access any CCB service. This means physically plugged into a building network port, or using a University VPN.
-
If you just want to use R or Jupyter, you can use our bespoke compute server with 128 CPUs and 2TB memory. For R, visit website https://rstudio.molbiol.ox.ac.uk ; for Jupyter, read our guide. Note that watchdogs manage resource use (see point #3).
-
To log into the CCB cluster with SSH, we have two login servers:
login1.molbiol.ox.ac.ukandlogin2.molbiol.ox.ac.uk. Use any SSH client to connect with a command such as:ssh user@login1.molbiol.ox.ac.uk. Both tmux and screen are installed for session persistence between logins. -
Login servers are provided for light use - file management, light development work, and submitting jobs to the cluster. As such, your login sessions are restricted to a small quantity of CPUs and memory similar to a personal computer, ensuring one user doesn't monopolise the system. For more computing resources, review our cluster documentation. As a last defense against becoming unresponsive, our login nodes and our R/Jupyter server run resource watchdogs - these prevent total resource use reaching 100%, which would effectively lock the system for everyone. See acceptable use and resource management for further details.
-
No graphical login is available. To learn more about the Linux command line, consider attending an OBDS training course. If you want to use RStudio or Jupyter, please consider using our services described above; for advanced users, you can use SSH tunnels with port forwarding, covered in our SSH tunnel guide. To locally view files on CCB, note that the
-Xoption to SSH is enabled for X forwarding, and that the image viewer feh and PDF viewer xpdf are both installed.
Security
Password
When your account was created you were provided with a randomly generated 16 character password.
You are welcome to keep this.
If you wish to change it, then login using SSH as described above and type the command passwd. Please note that:
-
what you type won't appear on the screen as you type it
-
your password must be at least 16 characters
-
it must not be a password used anywhere else
-
you must never, ever share it
For more information on choosing secure passwords, please see the University's Information Security website.
2FA
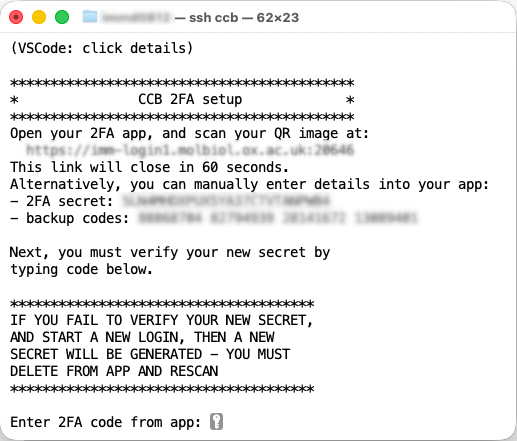 Use a plain terminal to setup & verify. MacOS: Terminal. Windows: PowerShell.
VSCode users: click
Use a plain terminal to setup & verify. MacOS: Terminal. Windows: PowerShell.
VSCode users: click details to open output tab.
You must immediately verify your new secret in the same login session, otherwise on next login a new secret is created.
On subsequent logins, your 2FA verification will be remembered for several hours before asking again.
To reset your 2FA we need another second-factor, either visit us inside WIMM or send us a photo of your university card with number clearly visible.
Customising your bashrc
Sooner or later you will be advised to change the contents of your .bashrc file - the master configuration for your shell that is automatically loaded on login.
Please be aware that a significant percentage of issues that we're asked to help with turn out to be the result of misconfiguration of a .bashrc and that you can even block your own logins. In particluar, we strongly advise against adding to your .bashrc commands that do any of following:
- load modules, Python venvs, or Conda
- alter your
LD_LIBRARY_PATH
If you find yourself considering this, please consider contacting us for help first. We may be able to suggest a better alternative and save you hours (or even days) of wasted time debugging.
Data storage and sharing
When your account is set up you are provided with a private personal /home/ directory with quota limit of 10GB.
this is intended for small but important files - configuration and settings, scripts, documents, source code etc.
For large files such as NGS data, or to share data with colleagues on the cluster, you can request collaboration projects, which exist in /project/.
Collaboration projects are a secure way for data to be shared with specific groups of people.
If a project owner, you just determine scope of data and the people it needs to be shared with,
then we create the project for you with access restricted to specified users.
Please note that the quota for a project is shared by all project members.
See our project documentation for more information.
To join an existing project, please ask the PI to contact us via help@imm.ox.ac.uk and ask for your username to be added.
You can additionally share data with other people over the internet using our public datashare service.
To share data via Globus, you need to install their CLI client then permit it to access your folders.
If you are handling data that needs especially high confidentially (e.g. human patient data) or where there are specific security requirements imposed by the data source or funder (e.g. UK Biobank data), please get in touch with us before uploading it.
Click for more information on file system layout & management.
Transferring files
Graphical
The graphical program we recommend is FileZilla.
To use FileZilla with our 2FA,
you need to create a Site with interactive login,
and disable multiple connections.
From the main toolbar, click File -> Site Manager..., then follow this video:
Click for video showing FileZilla setup
Once you have connected, the files in your home directory on CCB will be displayed in the lower right panel of FileZilla. These can be dragged to the left panel in order to copy files from the cluster to your local machine. Similarly, dragging files from the left panel to the right panel will copy them from your local machine to a directory on the cluster.
To avoid a password prompt on every operation, first consider setting up an SSH key.
If you still need FileZilla to remember your password:
Toolbar -> Edit -> Settings -> Interface -> Passwords -> check Save passwords with a master password
Command-line
If you are new to Linux, then we recommend starting with command-line program scp.
It behaves just like cp except it transfers files over SSH connections
e.g. scp file user@login1.molbiol.ox.ac.uk:/home/user/file
Once you gain experience, consider switching to program rsync.
This is more efficient than scp - skip files already transferred, and can resume an interrupted download.
More information
Software
A huge amount of time has been invested in providing access to hundreds of bioinformatics, Python and R packages "out-of-the-box". These are provided using the module system, which makes it quick and easy to swap between the different software packages and versions that you require. We recommend taking the time to read modules, python-cbrg and R-cbrg before attempting to run any programs or install software which appears to be "missing".
JADE also has the singularity container solution installed (aka apptainer), click here for more information.
Running jobs
The CCB cluster uses the Slurm job manager to allow you to run dozens (or even hundreds) of programs at the same time. The use of Slurm is too large a topic to cover in this introduction, so please take a look at slurm-basics for more details.
Databases
iGenomes is a collection of reference sequences and annotation files for commonly analysed organisms.
The files were originally generated by Illumina.
The files have been downloaded from Ensembl, NCBI, or UCSC, and chromosome names have been changed to be simple and consistent with their download source.
On the CCB servers these files can be found at /databank/igenomes/.
Click for more information
Staying up to date
Our change log maintains a running record of updates to JADE, and we also publicise known issues & limitations.
Further reading
We recommend that you take some time to read the entire documentation, especially the FAQ. Investing some time now can save many wasted hours trying to figure out how to do something later!