Storage
This page describes the file hierarchy on the JADE cluster.
At a high level, the various file locations are available across the whole cluster and you can read and write data from anywhere.
/ceph is the root mount of our primary data store and the place you should store most of your data.
A number of sub-locations within this are available via symbolic links described below.
Ceph overview
Our Ceph has 3-way replication - each file you store in Ceph has 3 copies. This is immediate and automatic whenever you create or modify a file, a built-in feature of Ceph. Two copies are co-located with the compute cluster in the data centre; the third copy is stored off-site in the WIMM building. This protects against:
- system faults, such as a storage disk failing or a copy being corrupted (bit rot)
- catastrophic disaster at one site
If Ceph detects corruption through its regular scanning, or a storage disk fails, then the other two copies enable automatic repair. This does NOT protect against user behaviour e.g. overwriting or accidental deletion - for this use our 24-hour backup (see below).
Further reading: Ceph documentation on replication and integrity
Key symbolic links
The / (root) filesystem on the CCB cluster contains a number of symbolic links which you can use to refer to other data locations.
We recommend using these rather than the absolute file paths where possible,
as we reserve the right to change the location of data where necessary but will endeavor to ensure that the links always point to the correct target.
/databank - A link to /ceph/databank/ and the location for reference genomes.
/home/[a-z] - A set of links to /ceph/home/[a-z] and the locations of home directories.
/miseq - A link to /ceph/miseq and the location where new NGS data is uploaded.
/project - A link to /ceph/project/ and the location for collaboration projects.
/package - A link to /ceph/package/OS_VERSION which contains the computational biology software which is provided through our module system.
Quota
Managing quota
Run our tool getquota to output summary of your quota use on CCB file systems.
The output will resemble the following:
$ getquota
===============
HOME QUOTA (GB)
===============
/home/a/aowenson quota=10.0 size=3.0 usage=30.0%
===================
PROJECT QUOTAS (TB)
===================
/project/sysadmin quota=10.0 size=2.0 usage=20.0%
Your home quota is for you alone, whereas the project quotas and the usage figures are for the entire project and all users in it.
To list usages of home & project contents, append argument -l:
$ getquota -l
===============
HOME QUOTA (GB)
===============
QUOTA SIZE USAGE
/home/a/aowenson 10.0 3.6 36.0%
.cache 1.3 13.0%
.local 0.2 2.0%
.vscode-server 1.3 13.0%
...
===================
PROJECT QUOTAS (TB)
===================
QUOTA SIZE USAGE
/project/sysadmin 10.0 2.0 20.0%
aowenson 0.1 1.0%
datashare 0 0
shared 1.7 17.0%
getquota is simply a wrapper around our tool cephls,
so for deeper usage analysis you can use this e.g cephls -lt /project/sysadmin/shared.
Exceeding your quota
If you exceed your home quota you will not be able to work; you will have to delete some data or move it to a collaboration project. We do not offer additional home quota. If anyone in a secure collaboration project exceeds the quota then everyone in the project will lose the ability to create more data and your PI will need to pay for more. See accounts for more details on how to do this.
Folder ~/.cache is safe to delete.
But do NOT delete ~/.local without fully understanding its contents, this is where your programs and applications store libraries and state such as RStudio.
If you cannot delete an item and think you cannot move it, you can use a symlink to move the data but have it still appear in your home.
But the risk is if you lose project access, you lose access to the data.
Example: move your .local folder to a project (stop your applications first e.g. RStudio):
$ mv ~/.local /project/sysadmin/aowenson/
$ ln -s /project/sysadmin/aowenson/.local ~/.local
$ ls -l ~/.local
/home/a/aowenson/.local -> /project/sysadmin/aowenson/.local
Transfer data
rsync
Command-line program rsync is more powerful than program scp, and so has many more options.
Useful features for you are: skip files already transferred, and can resume an interrupted download.
Useful arguments:
--recursive, -r you almost-always want this, unless auto-set by another argument
--archive, -a = -rlptgoD = recursive, preserve most metadata
--checksum, -c use file checksum hash instead of file size/timestamp to identify changes
--partial let transfer be resumed
--append-verify resume an interruped transfer
--compress, -z compress data during transfer
--progress show progress
--verbose, -v print each filename transferred
Typical tasks:
- transfer a folder:
rsync -a --progress ... - resume interrupted transfer:
rsync -a --progress --partial --append-verify ...
sftp
Downloading data from a collaborator using sftp is interactive and slightly different than using scp/rsync:
it downloads to where you started the program, instead of you specifiying location in an argument.
Example:
# Prepare download location
$ mkdir -p /project/sysadmin/Data/FromCollaborator
# Start inside download location
$ cd /project/sysadmin/Data/FromCollaborator
# Start sftp session
sftp user@collaborator.com # <- do not type sftp://
$ user@collaborator.com's password: # <- manually type/paste password
Connected to collaborator.com.
sftp>
# Browse with usual commands: cd, ls, pwd:
sftp> ls
10-123456
sftp> cd 10-123456
sftp> pwd
Remote working directory: /10-123456
sftp> ls
data report.html
sftp> cd ..
sftp> pwd
Remote working directory: /
# Download 1 file
sftp> get 10-123456/report.html
Fetching /10-123456/report.html to report.html
...
# Download 1 folder
sftp> get -r 10-123456
Fetching /10-123456/ to 10-123456
...
# Exit & review
sftp> exit
$ pwd
/project/sysadmin/Data/FromCollaborator
$ ls
10-123456
samba
This is specifically for sending to/from a Samba share outside CCB.
You will use program rclone https://rclone.org, as this does not involve mounting the Samba share.
First, create record of your target Samba in file ~/.config/rclone/rclone.conf e.g.:
[RFS]
type = smb
host = connect.ox.ac.uk
Load your Samba credentials:
echo "Enter your password:"
read -s PW
export RCLONE_SMB_USER='your_SSO'
export RCLONE_SMB_PASS="$(rclone obscure "$PW")"
export RCLONE_SMB_DOMAIN='ad.oak.ox.ac.uk'
Connect and trasfer:
rclone ls "$HOST:$SHARE_NAME"
rclone copy --progress local-file "$HOST:$SHARE_NAME"
rclone copy --progress "$HOST:$SHARE_NAME/remote-file" ./
Recovering project data
You can recover an accidentally deleted file, or revert a change, if done during the same day. A 24-hour snapshot is maintained for each project, the state of its data from yesterday. This is updated overnight each night, so you can only recover to yesterday.
The snapshot is stored in: /project/PROJECT-NAME/.snap/yesterday/.
Just browse for the file/folder and copy e.g.:
# Accidental delete:
$ rm -r /project/sysadmin/aowenson/Debug/apptainer-gpu
# Recover
$ cp -ar /project/sysadmin/.snap/yesterday/aowenson/Debug/apptainer-gpu ./
To avoid accidental deletion you can make your own files read-only: chmod u-w path/to/file .
As owner you can reverse at any time: chmod u+w path/to/file .
Duplicate files scan
In the background we are scanning your projects for duplicate files, currently every 28 days.
The last duplicate scan report can be viewed by running: getdups project-name.
Output will be a list of duplicated file groups e.g.:
...
"File hash = ccf0a698...": [
"File size = 10.2 GB",
"/ceph/project/sysadmin/aowenson/Images/CentOS.iso",
"/ceph/project/sysadmin/aowenson/Sandbox/image/CentOS.iso",
]
}
"108.5 GB can be reclaimed by deleting duplicated files"
Last duplicate scan was: 2024-09-07 05:13
The file hash can be considered a unique identifier of a files' contents, as the chance of a false positive are extremely remote - 1 in 2^256. So just exercise a little caution before deleting files, a false positive should be obvious.
You could resolve a duplication by deleting all-but-one of the files, but this could break your work.
Another resolution is creating a symbolic link - a pointer to the real file.
The command is: ln -s TARGET-FILE LINK-NAME.
E.g. to resolve the above duplication, I could run:
$ ln -s /project/sysadmin/aowenson/Images/CentOS.iso /project/sysadmin/aowenson/Sandbox/image/CentOS.iso
$ ls -l /project/sysadmin/aowenson/Sandbox/image/CentOS.iso
... /project/sysadmin/aowenson/Sandbox/image/CentOS.iso -> /project/sysadmin/aowenson/Images/CentOS.iso
To learn more about the scan, you can inspect code of getdups as it is simply a Bash script: less /usr/local/bin/getdups.
The raw scan data is stored at: /project/PROJECT-NAME/.usage/PROJECT-NAME-rmlint-1G.json.
This is generated by tool rmlint.
Project files explorer
Using our Jupyter Notebook server, we have developed a Python tool to interactively and visually inspect your project quota use.
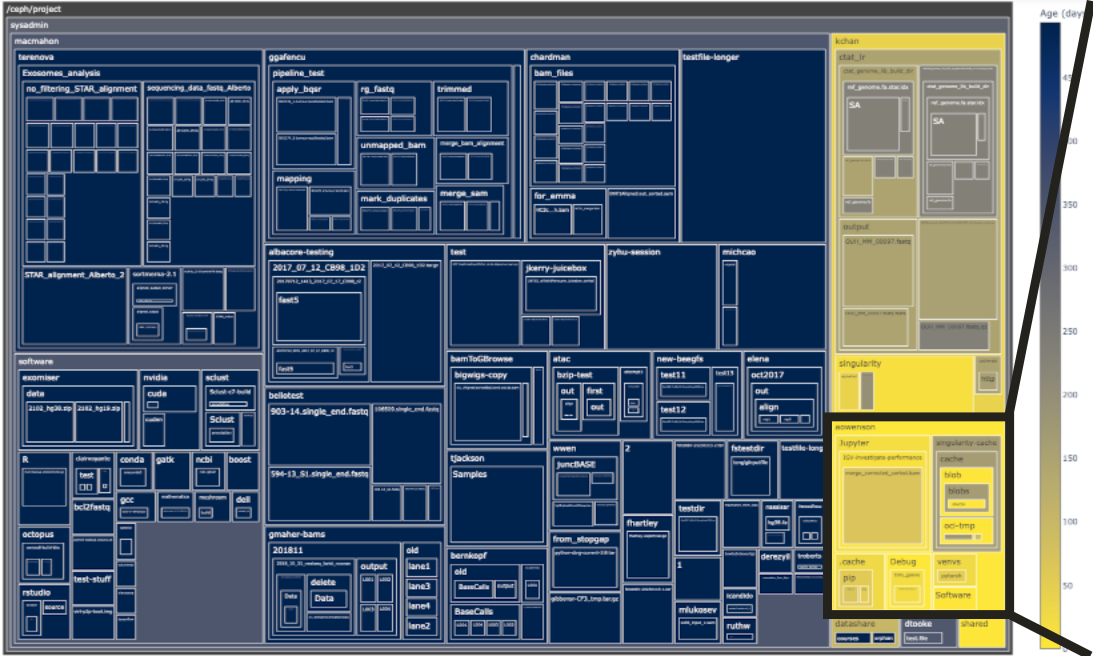
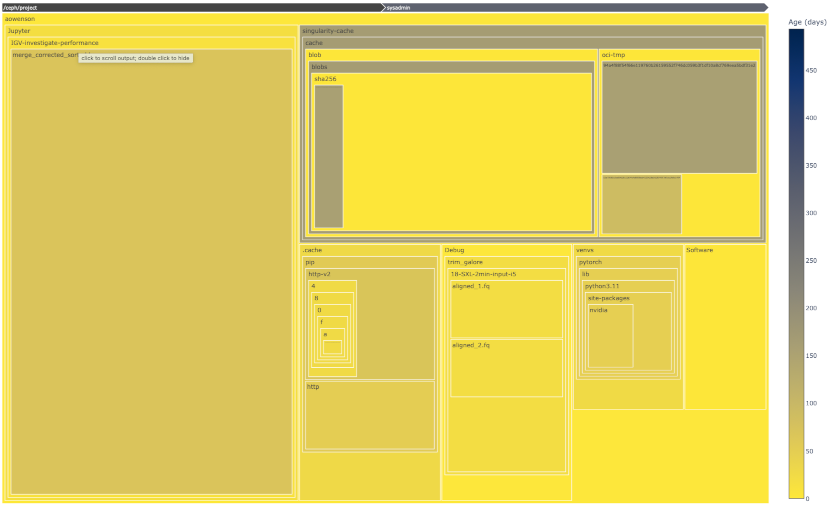
Box area represents disk size, colour represents age (yellow=young blue=old), and clicking a box expands into it.
Once in our Jupyter Notebook server, simply run this Python code to get started:
from ccb_file_utils import VisualiseUsage
v = VisualiseUsage('/project/PROJECT-NAME')
v.view()
This uses our custom folder scanner, which avoids scanning small folders to complete the scan much faster, and a database cache to avoid unnecessary rescans. Default threshold is 0.1% of project size, this should complete a scan in 15-30 seconds.
To access raw scan data as a Pandas DataFrame:
df = v.df
To customise the scan or treemap, inspect the documentation:
help(VisualiseUsage)
Differences from t1-data
The CCB historically used storage referred to as t1-data which is now retired.
For long-time CCB users, there are some key differences:
-
There are no users quotas for the Ceph; all quotas are per-project
-
File paths which previously started
/t1-datano longer work -
All data is stored in 3 copies at 2 separate physical locations for resilience; there are no separate backups and the old
/t1-databackups have been switched off -
To find out how much data a directory contains, use command
ls -ldh path/to/directory. This is much faster thanduthanks to a special feature in Ceph, but please be aware that this total may take 1-2 hours to update when data is added or removed.